
 University of Nottingham
University of NottinghamI am a Ph.D. candidate at University of Nottingham, specializing in Computer Science and Statistics. My research interests lie in computer vision, computational pathology, and multi-modal large language models, with a focus on developing intelligent systems for medical image analysis and cross-modal understanding.
Before starting my Ph.D., I worked as a Research Assistant at Westlake University in Yang Lin's Laboratory. I received my M.Sc. degree in Data Science from the University of Glasgow and my B.Eng. degree in Computer Science and Technology from Qufu Normal University.
Warning
Problem: The current name of your GitHub Pages repository ("Solution: Please consider renaming the repository to "
http://".
However, if the current repository name is intended, you can ignore this message by removing "{% include widgets/debug_repo_name.html %}" in index.html.
Action required
Problem: The current root path of this site is "baseurl ("_config.yml.
Solution: Please set the
baseurl in _config.yml to "Education
-
 University of NottinghamPh.D. in Computer Science and Statistics2024 - present
University of NottinghamPh.D. in Computer Science and Statistics2024 - present -
 University of GlasgowM.Sc. in Data Science2021 - 2022
University of GlasgowM.Sc. in Data Science2021 - 2022 -
 Qufu Normal UniversityB.Eng. in Computer Science and Technology2016 - 2020
Qufu Normal UniversityB.Eng. in Computer Science and Technology2016 - 2020
Experience
-
 Westlake UniversityResearch Assistant2022 - 2024
Westlake UniversityResearch Assistant2022 - 2024 -
 Dipath Technology Co., Ltd.Algorithm Engineer2022 - 2024
Dipath Technology Co., Ltd.Algorithm Engineer2022 - 2024 -
 The Third Affiliated Hospital of Sun Yat-sen UniversityResearch Collaborator, Department of Pathology2022 - 2023
The Third Affiliated Hospital of Sun Yat-sen UniversityResearch Collaborator, Department of Pathology2022 - 2023
News
Selected Publications (view all )
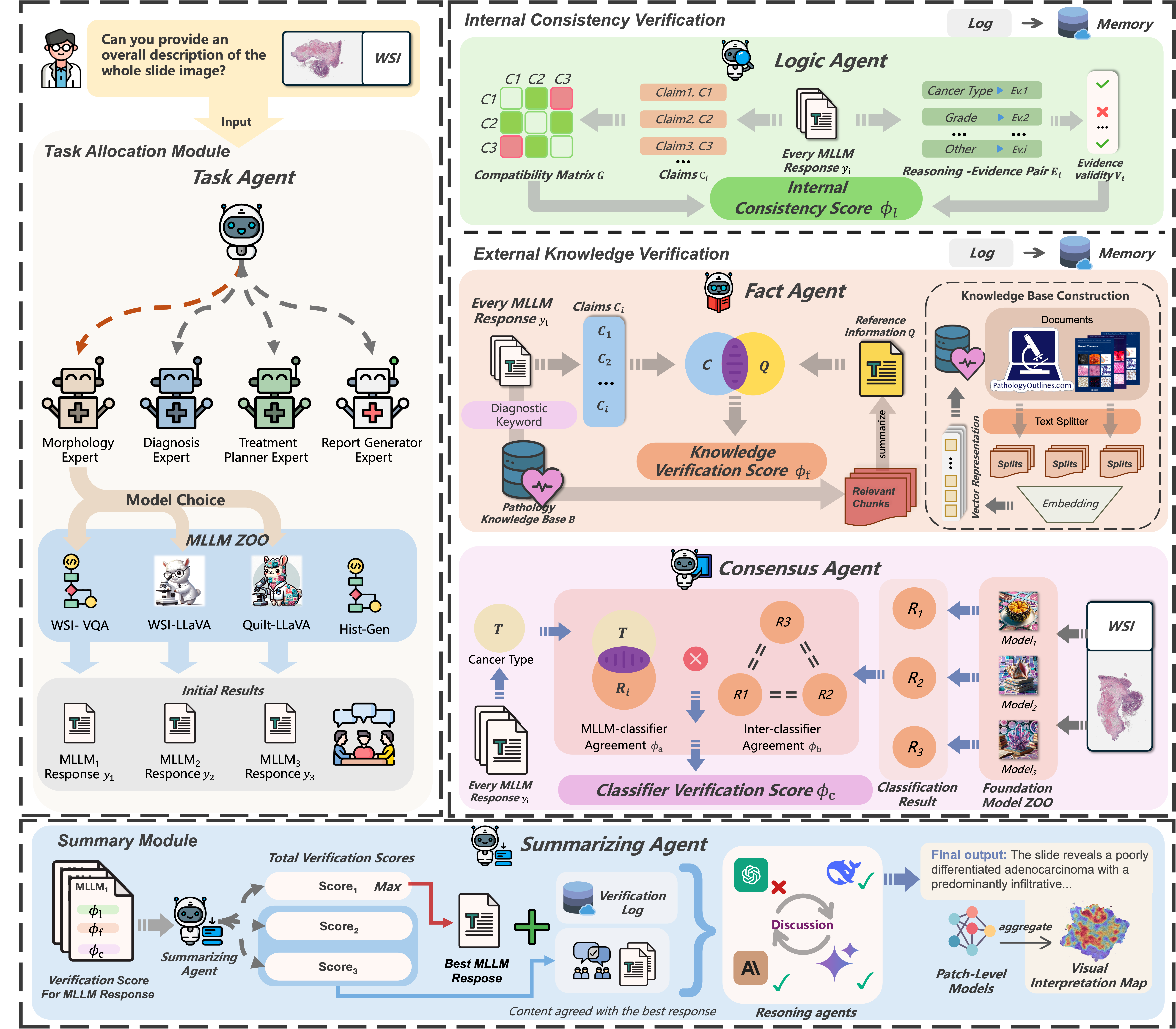
WSI-Agents: A Collaborative Multi-Agent System for Multi-Modal Whole Slide Image Analysis
Xinheng Lyu, Yuci Liang, Wenting Chen, Meidan Ding, Jiaqi Yang, Guolin Huang, Daokun Zhang, Xiangjian He, Linlin Shen
International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI) 2025 Oral
We propose WSI-Agents, a novel collaborative multi-agent system for multi-modal WSI analysis that integrates specialized functional agents with robust task allocation and verification mechanisms. The system enhances both task-specific accuracy and multi-task versatility through three key components: task allocation, verification mechanisms, and summary modules.
WSI-Agents: A Collaborative Multi-Agent System for Multi-Modal Whole Slide Image Analysis
Xinheng Lyu, Yuci Liang, Wenting Chen, Meidan Ding, Jiaqi Yang, Guolin Huang, Daokun Zhang, Xiangjian He, Linlin Shen
International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI) 2025 Oral
We propose WSI-Agents, a novel collaborative multi-agent system for multi-modal WSI analysis that integrates specialized functional agents with robust task allocation and verification mechanisms. The system enhances both task-specific accuracy and multi-task versatility through three key components: task allocation, verification mechanisms, and summary modules.
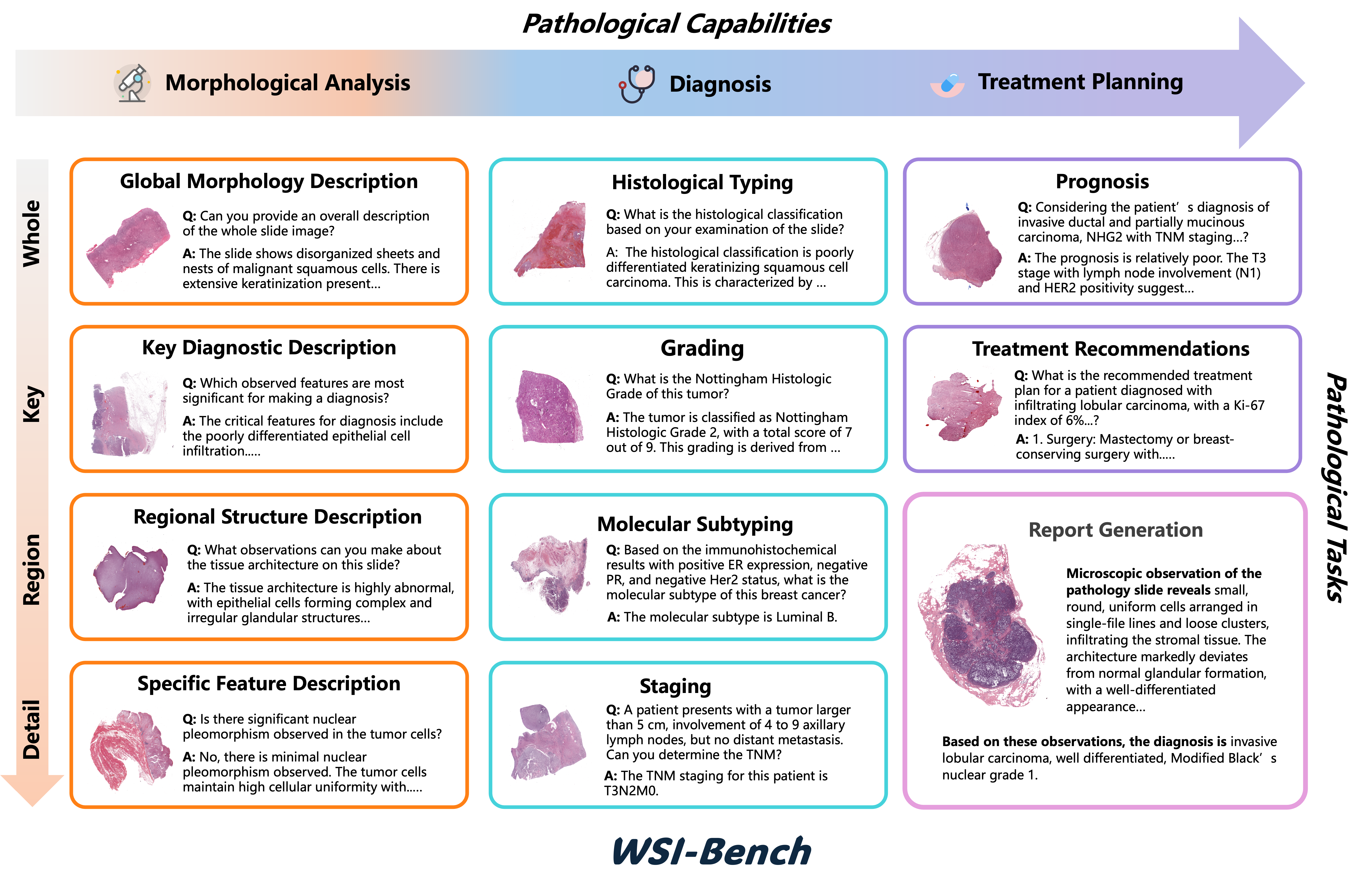
WSI-LLaVA: A Multimodal Large Language Model for Whole Slide Image
Yuci Liang*, Xinheng Lyu*, Wenting Chen, Meidan Ding, Jipeng Zhang, Xiangjian He, Song Wu, Xiaohan Xing, Sen Yang, Xiyue Wang, Linlin Shen (* equal contribution)
International Conference on Computer Vision (ICCV) 2025
We introduce WSI-LLaVA, an MLLM framework for gigapixel WSI understanding with a three-stage training strategy that provides detailed morphological findings to explain diagnostic reasoning. We also present WSI-Bench, the first large-scale morphology-aware benchmark containing 180k VQA pairs from 9,850 WSIs across 30 cancer types.
WSI-LLaVA: A Multimodal Large Language Model for Whole Slide Image
Yuci Liang*, Xinheng Lyu*, Wenting Chen, Meidan Ding, Jipeng Zhang, Xiangjian He, Song Wu, Xiaohan Xing, Sen Yang, Xiyue Wang, Linlin Shen (* equal contribution)
International Conference on Computer Vision (ICCV) 2025
We introduce WSI-LLaVA, an MLLM framework for gigapixel WSI understanding with a three-stage training strategy that provides detailed morphological findings to explain diagnostic reasoning. We also present WSI-Bench, the first large-scale morphology-aware benchmark containing 180k VQA pairs from 9,850 WSIs across 30 cancer types.